MOBscan
MOBscan is a web application for identifying relaxase MOB families. It uses the hmmscan function of the HMMER3 software suite (Eddy, 2011) to search against MOBfamDB, a curated relaxase profile HMM database. If you find it useful for your work, please cite it as:
| Garcillán-Barcia M.P., Redondo-Salvo S., Vielva L., de la Cruz F. (2020) "MOBscan: Automated Annotation of MOB Relaxases". In: de la Cruz F. (eds) Horizontal Gene Transfer. Methods in Molecular Biology, vol 2075. Humana, New York, NY |
MOBfamDB construction information
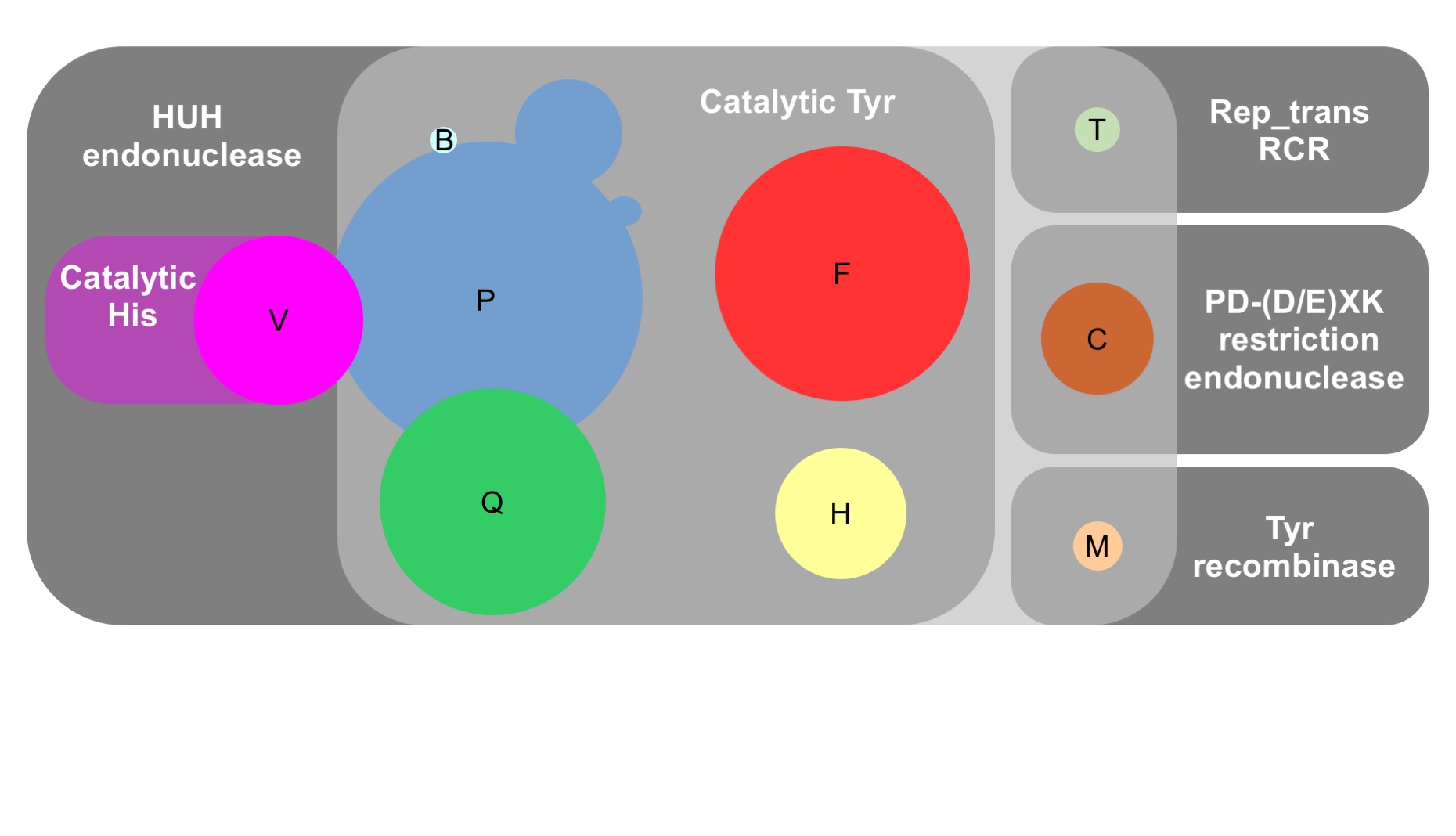
Individual profile HMM files for relaxase families MOBP (T4SS_MOBP1, T4SS_MOBP2, and T4SS_MOBP3), MOBQ (T4SS_MOBQ), MOBH (T4SS_MOBH), MOBC (T4SS_MOBC), MOBV (T4SS_MOBV), and MOBB (T4SS_MOBB) have been retrieved from the TXSScan suite (Abby et al., 2016).
New profile HMMs were built for MOBF, MOBT, and MOBM families from seed multiple sequence alignments generated with MUSCLE v3.8.31 (Edgar, 2004) by using the hmmbuild function of HMMER3.
MOBF Profile
The profile HMM for the MOBF relaxase family (profile_MOBF) was generated from an alignment of the N-terminal 300 residues of 146 MOBF proteins described by (Garcillan-Barcia et al., 2009), which was previously trimmed to remove poorly aligned regions at the edges by using Trimal (Capella-Gutierrez et al., 2009).
MOBT Profile
The profile HMM for the MOBT relaxase family (profile_MOBT) was built from an alignment of 498 relaxases retrieved from a BLASTP search (Altschul et al., 1990) using as query the Orf20_Tn916 complete sequence present in Figure 3 of (Wright and Grossman, 2016). The alignment was trimmed while keeping both HTH and Rep_trans domains.
MOBM Profile
For the novel family MOBM, a profile HMM (profile_MOBM) was built based on an alignment of thirteen relaxases from Clostridium perfringens plasmids, retrieved from a BLASTP search using TcpM_pCW3 (GenBank Acc. No. WP_003479716.1) as query.